Load GW Data
Load-data.Rmd
library(beacon)
#>
#> Attaching package: 'beacon'
#> The following object is masked from 'package:utils':
#>
#> vi
#> The following objects are masked from 'package:base':
#>
#> mode, pipe1. Download GW data from GWOSC
beacon employs GWOSC API (v2) to download the public GW
data. Therefore, you need the network connection.
There are helper functions, such as,
list_gwosc_event([##(1) Check available GW events])list_gwosc_param([##(3) Check parameters])
to check the right input for
download_*/get_* functions:
download_event([##(2) Download event data])get_gwosc_param([##(3) Check parameters])
(1) Check available GW events
GW_event_list <- list_gwosc_event()
cat("First 6 events:\n")
#> First 6 events:
print(head(GW_event_list))
#> [1] "GW240109_050431" "GW240107_013215" "GW240105_151143" "GW240104_164932"
#> [5] "GW231231_154016" "GW231231_120147"(2) Download event data
Here we test with GW150914. Download event data using
download_event().
download_event(event_name = "GW150914", # event name
det = "H1", # Detector name
path = "test_data/", # download path
file.format = 'hdf5', # default
sampling.freq = 4096 # default
)
# YOU'LL SEE:
#
# > Trying event-level strain-files: https://gwosc.org/api/v2 /events/GW150914/strain-files
# -> GET https://gwosc.org/api/v2/events/GW150914/strain-files
# > Downloading: https://gwosc.org/archive/data/O1/1126170624/H -H1_LOSC_4_V1-1126256640-4096.hdf5
# -> to: test_data//H-H1_LOSC_4_V1-1126256640-4096.hdf5
# trying URL 'https://gwosc.org/archive/data/O1/1126170624/H -H1_LOSC_4_V1-1126256640-4096.hdf5'
# Content type 'application/octet-stream' length 130159479 bytes (124.1 MB)
# ==================================================
# downloaded 124.1 MBIf you want to:
- analyze data directly, and
- save your storage.
Set load=TRUE, and remove=TRUE . In this
case you don’t need to specify path (default path is
“./”).
(3) Check parameters
For available parameters, you can check by
param_names <- list_gwosc_param()
cat("First 6 parameters:\n")
#> First 6 parameters:
print(head(param_names))
#> [1] "GPS" "mass_1_source" "mass_1_source_lower"
#> [4] "mass_1_source_upper" "mass_1_source_unit" "mass_2_source"Then collect parameter value(s) of specific event(s) as follows.
One parameter for an event:
params <- get_gwosc_param(
name = 'GW150914',
param = 'GPS'
)
cat("GPS time of GW150914:\n")
print(params) # returns 1x1 data.frame
# GPS time of GW150914:
# GPS
# GW150914 1126259462Multiple parameters for an event:
params <- get_gwosc_param(
name = 'GW150914',
param = c('mass_1_source',
'mass_2_source',
'luminosity_distance',
'GPS')
)
cat("Some parameters of GW150914:\n")
print(params) # returns 1x4 data.frame
# Some parameters of GW150914:
# mass_1_source mass_2_source luminosity_distance GPS
# GW150914 35.6 30.6 440 1126259462Or just all the available parameters for an event:
params <- get_gwosc_param(
name = 'GW150914',
param = 'all'
)
print(params) # returns 1x45 data.frame
#
# GPS mass_1_source mass_1_source_lower mass_1_source_upper mass_1_source_unit ...
# GW150914 1126259462 35.6 3.1 4.7 M_sun ...
# 2. Load HDF5 Data
Since R doesn’t have any package for reading gwf files
for the moment, we mainly treat hdf5 (or h5)
file format.
Using following functions, one can read and write hdf5
files:
read_H5write_H5
Basically those functions refer to the structure of hdf5
as follows.
meta/GPSstart: Starting GPS time of strain data is stored.strain/Strain: Strain data is stored.
Additionally, if GW data has a Data Quality (DQ), it will refer to:
-
quality/simple/DQmask: DQ mask of bits is stored in decimal unit.
(1) Read hdf5 file
For using simluated demo data for the illustration, it can be found in following directory:
demo.file <- system.file("extdata", "demo_4kHz.h5", package = "beacon")
cat('Demo hdf5 data file directory:\n')
#> Demo hdf5 data file directory:
print(demo.file)
#> [1] "/home/runner/work/_temp/Library/beacon/extdata/demo_4kHz.h5"(The demo data was created by using Python’s pycbc
package)
read_H5 will load strain data and GPS start time to
construct ts object (here, you need to know the sampling
frequency, e.g. 4096 Hz).
demo <- read_H5(
file = demo.file,
sampling.freq = 4096,
dq.level = 'all' # defulat: "BURST_CAT2"
)
cat(
"Demo Data Summary:\n",
"- Starting time:", ti(demo), "s\n",
"- End time:", tf(demo), "s\n",
"- Duration:", tl(demo), "s\n",
"- Time range:", paste(tr(demo), collapse=' - '), "s\n",
"- Sampling frequency:", frequency(demo), "Hz\n"
)
#> Demo Data Summary:
#> - Starting time: 0 s
#> - End time: 31.99976 s
#> - Duration: 32 s
#> - Time range: 0 - 31.999755859375 s
#> - Sampling frequency: 4096 Hz
print(demo)
#> Time Series:
#> Start = c(0, 1)
#> End = c(31, 4096)
#> Frequency = 4096
#> [1] -1.402802e-20 -1.451734e-20 -1.529089e-20 -1.461478e-20 -1.516181e-20
#> [6] -1.447528e-20 -1.515767e-20 -1.474572e-20 -1.496885e-20 -1.535953e-20
#> [11] -1.498296e-20 -1.472902e-20 -1.564059e-20 -1.546743e-20 -1.471808e-20
#> [16] -1.587687e-20 -1.512821e-20 -1.579603e-20 -1.493327e-20 -1.541508e-20
#> [ reached 'max' / getOption("max.print") -- omitted 131052 entries ]
#> attr(,"dqmask")
#> DATA CBC_CAT1 CBC_CAT2 CBC_CAT3 BURST_CAT1 BURST_CAT2 BURST_CAT3
#> [1,] 1 1 1 1 1 1 1
#> [2,] 1 1 1 1 1 1 1
#> [ reached 'max' / getOption("max.print") -- omitted 30 rows ]
#> attr(,"dqmask")attr(,"tsp")
#> [1] 0 31 1
#> attr(,"dqmask")attr(,"level")
#> [1] allPlot oscillogram:
plot_oscillo(demo)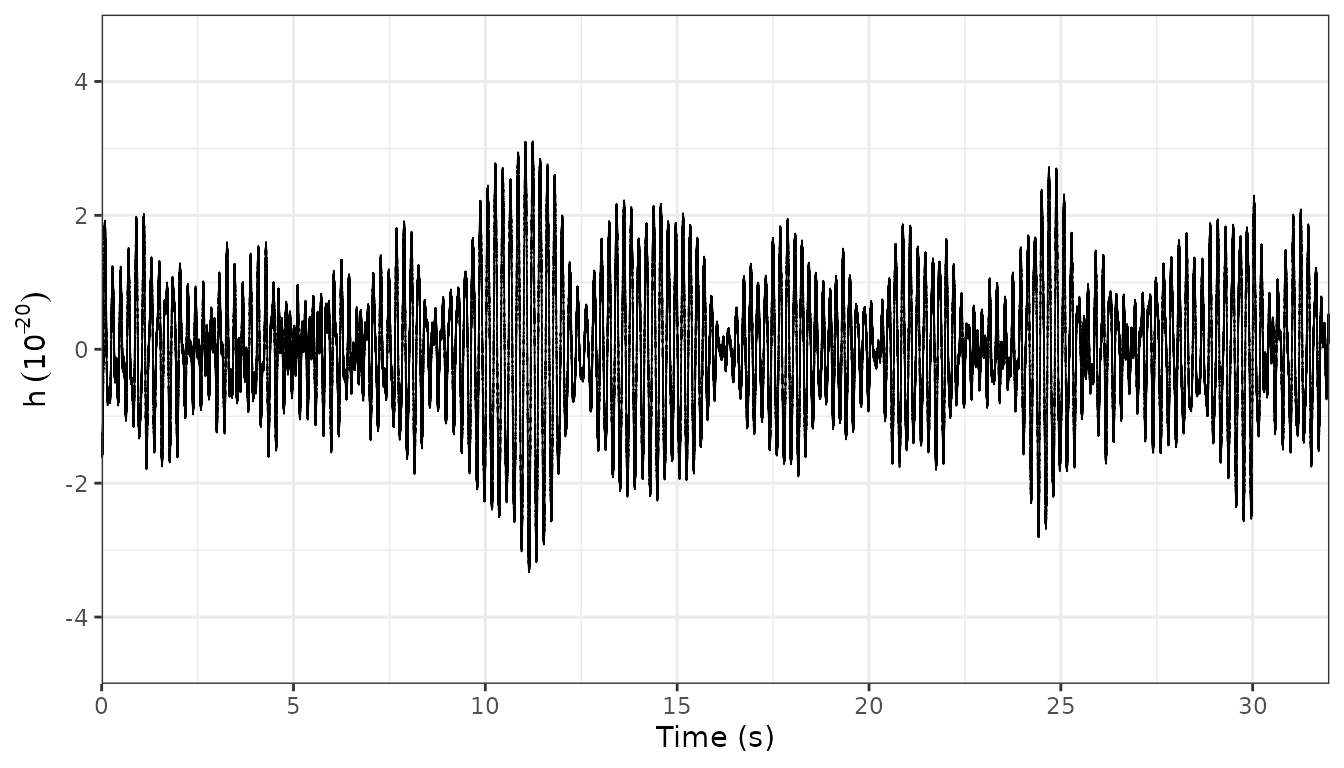
Using get_dqmask, you can extract the (multivariate)
time series of DQ bits with the sampling frequency of 1 Hz.
print(get_dqmask(demo))
#> Time Series:
#> Start = 0
#> End = 31
#> Frequency = 1
#> DATA CBC_CAT1 CBC_CAT2 CBC_CAT3 BURST_CAT1 BURST_CAT2 BURST_CAT3
#> 0 1 1 1 1 1 1 1
#> 1 1 1 1 1 1 1 1
#> [ reached 'max' / getOption("max.print") -- omitted 30 rows ]
#> attr(,"level")
#> [1] allThis will be used in DQ veto of BEACON pipeline.
(2) Write hdf5 file
Similarly to read_H5, write_H5 also refers
to basic directories of: meta/GPSstart, and
strain/Strain.
If you put more meta info, you can follow the code below:
write_H5(
file = 'path/to/your/hdf5/file.h5',
demo,
meta.list =
list(
'meta/sample_freq' = frequency(demo),
'quality/simple/DQmask' = rep(127, # All 1 of bit
32) # 32 s data case
)
)⚠️ Note: In current version,
read_H5does not read all the meta information in hdf5 file’s directory.